Tutorial
Click to download Tutorial.pdf
Within the DBPOM, we have developed a tool for distinguishing the reversed and adverse genes of a specified compound, or drug on cancer samples, by comparing the data stemming from the two groups of differential gene expression results.
Differential gene expression analyses have been performed on both cancer versus control tissues samples (E1) from TCGA and molecular-compound-treated cell lines versus DMSO-treated cell lines (E2) from the CMap database. When a cell line is treated with a compound, if the effect on a gene is oppositional, meaning it is up-regulated in E1 and down-regulated in E2 or vice versa, we consider it to have a reversed effect on said gene. Otherwise, if a gene is marked as 1(E1) ~1(E2), -1(E1) ~-1(E2), 0(E1) ~1(E2) or 0(E1) ~-1(E2), with 0 standing for no change, we consider the compound to have an adverse effect on this specific gene. For each compound, if it reverses the expression of a cancer-associated gene, it is thought to be effective on the gene. On the contrary, if the gene expression substantially deviates after treatment with the compound, we believe that the drug has had an adverse effect on the gene. Moreover, we defined one score as the Reversed and Adverse of Drug Effect (RADE) which indicates the safety and effectiveness of each compound by integrating both the reversed and adverse genes.
Regarding precision medicine, we consider two commonly studied cases. One is for specific mutations, as users may want to know the differences of drug response by individual patients with vs. without a specific mutation; and the other is about the overall mutation pattern as different cell lines of the same cancer type may respond very differently to the same drug due to their distinct mutation patterns. We have developed a tool for measuring the similarity between the mutation profiles in a cell line and in the cancer of a patient. For each cancer sample, the cell line with the highest similarity score was considered as the best reference.
There are four main modules in DBPOM.
1. Compounds
User could get the drug response by inputting the cancer type, cell line ID, and compound name.
The datasets provide three options: (1) TCGA vs. CMap (2) subTCGA vs. CMap (all mutaion) (3) subTCGA vs. CMap (specific mutation):
(1) TCGA versus CMap: in E1, all of the cancer samples of the selected cancer type in TCGA are utilized. While this is the general strategy in current research, it does not consider individual differences. We provide this option in case some users need it.
(2) subTCGA-mutaion versus CMap: in E1, the subset of cancer samples in TCGA which have high matching scores with selected cell line are utilized. We provide this strategy as overall mutation pattern is an important criterion for distinguishing different cell lines. So, we provide users with this option to suit diverse requirements. In practice, for each cancer patient, he or she is firstly classified into the most suitable sample group according to mutation similarity, and then to analyze effect of drugs on him or her.
(3) subTCGA-specific versus CMap: E1 utilizes a subset of cancer samples with the specific mutation that user concerns from TCGA, which match the specific mutation that user concerns of the selected cell line in E2. We proposed this strategy and expected to get a reasonable evaluation of drug effectiveness that could then assist precision medicine.
Example: BRCA, BT20, erlotinib and subTCGA(specific mutation) vs. CMap
By clicking Filter, the search result will be shown as followings.
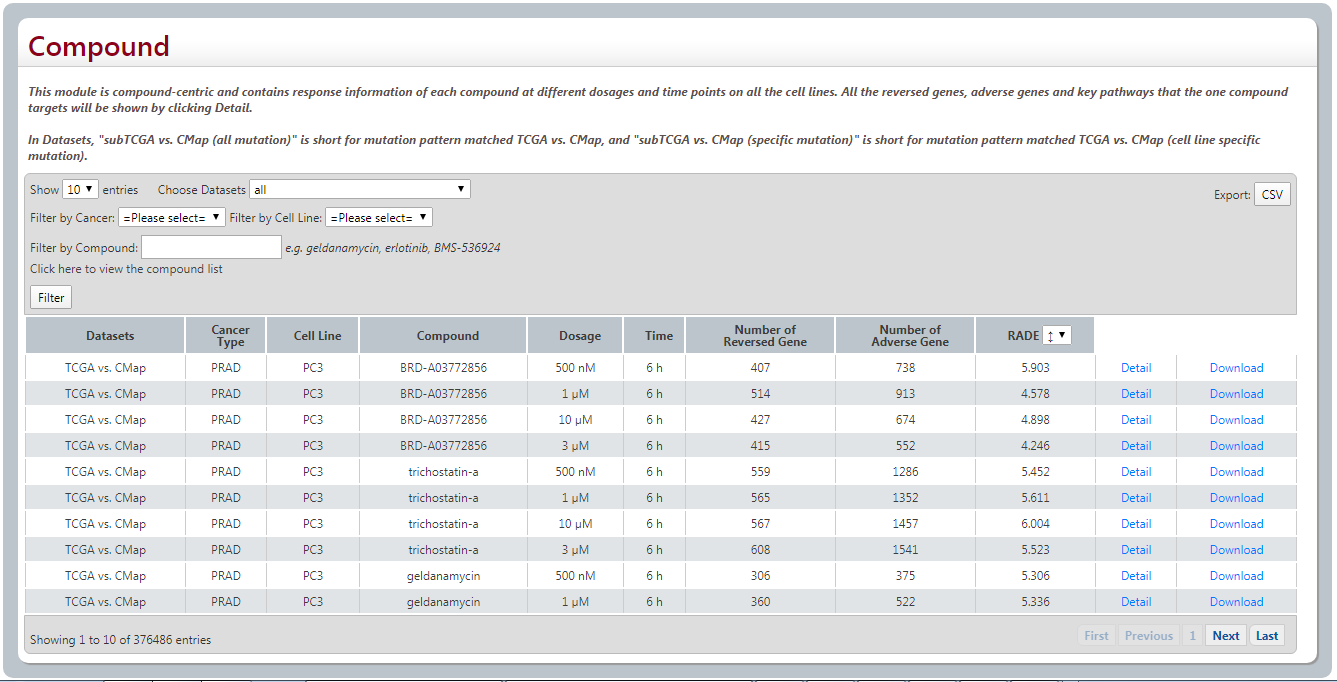
“Number of Reversed gene” is the gene number of reversed genes. “Number of Adverse gene” is the gene number of the adverse genes. “RADE” is a score defined by considering both the number of reversed genes and adverse genes. The lower the score, the higher the effectiveness and safety.
Users could download all the reversed genes and adverse genes, and their differentially expressed level in E1(cancer vs. normal) and E2(compound treated cell line vs. DMSO treated cell line) by clicking download.
By clicking detail, it will jump to compound detail page.
(1) Overview
User could search the detail information of this compound or drug from CMap by clicking “Jump”. For some drugs that reported in GDSC, the information is also provided in our page.
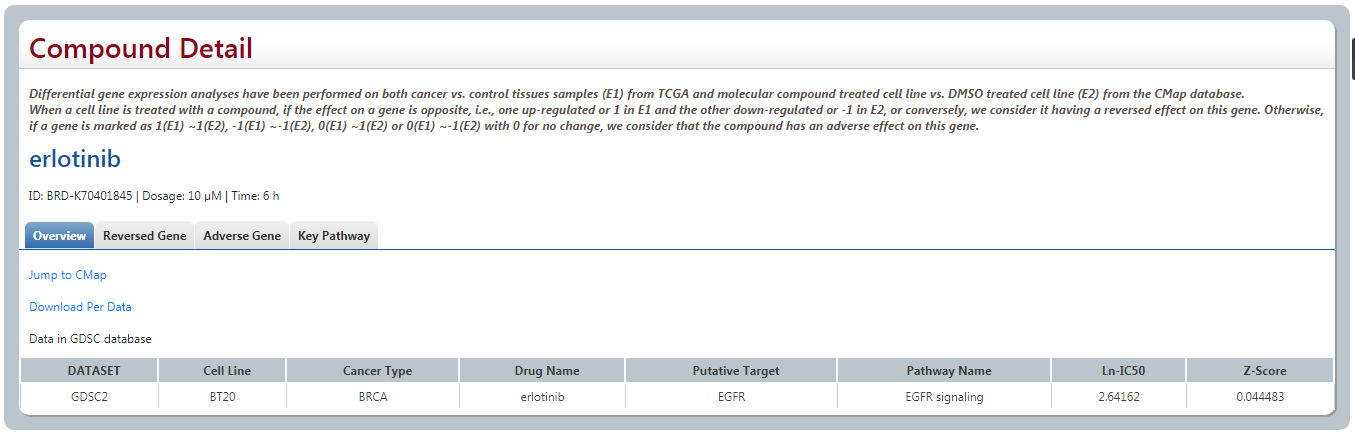
(2) Reversed gene
By clicking Reversed Gene, it will list all the reversed genes with differential expression level in E1(cancer vs. normal) and E2(compound treated cell line vs. DMSO treated cell line). User could sort the value by clicking arrows. We provide Download in three formats CSV, Excel and PDF.
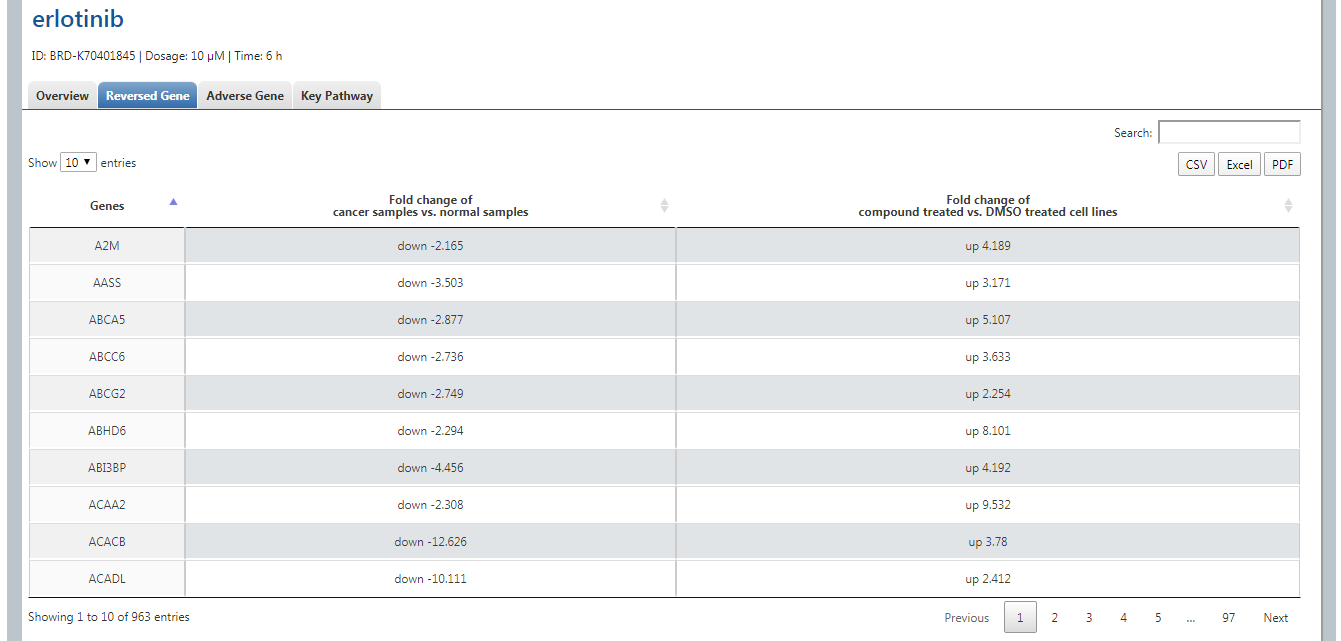
Four gene function annotation methods (GO, KEGG, PID, REACTOME) are provided by clicking the relative buttons. The pathways with pvalue < 0.05 are the significant ones.
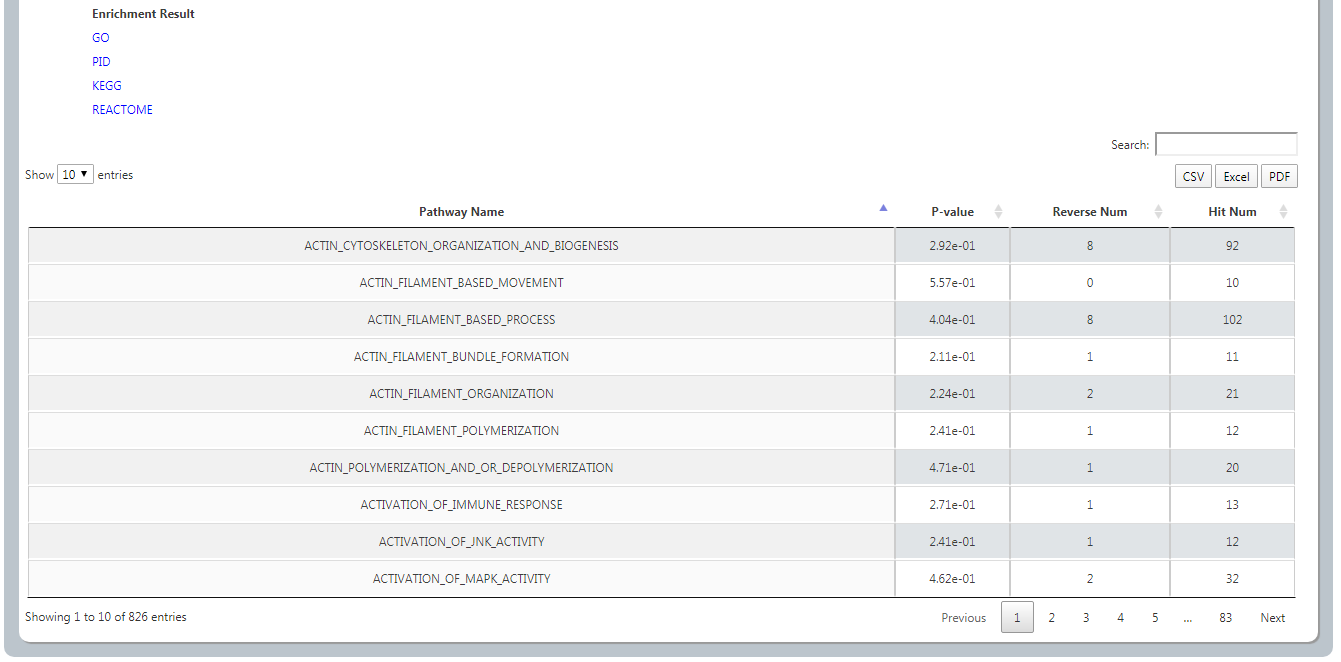
(3) In a similar way, analyses of the adverse gene are following the same way.
(4) We additionally provide 11 key pathways and their sub pathways that clinicians and drug researchers may pay much more attention.
When user selects “Cell cycle” and then “KEGG_CELL_CYCLE”, and clicks Submit, all the reversed and adverse genes are shown as
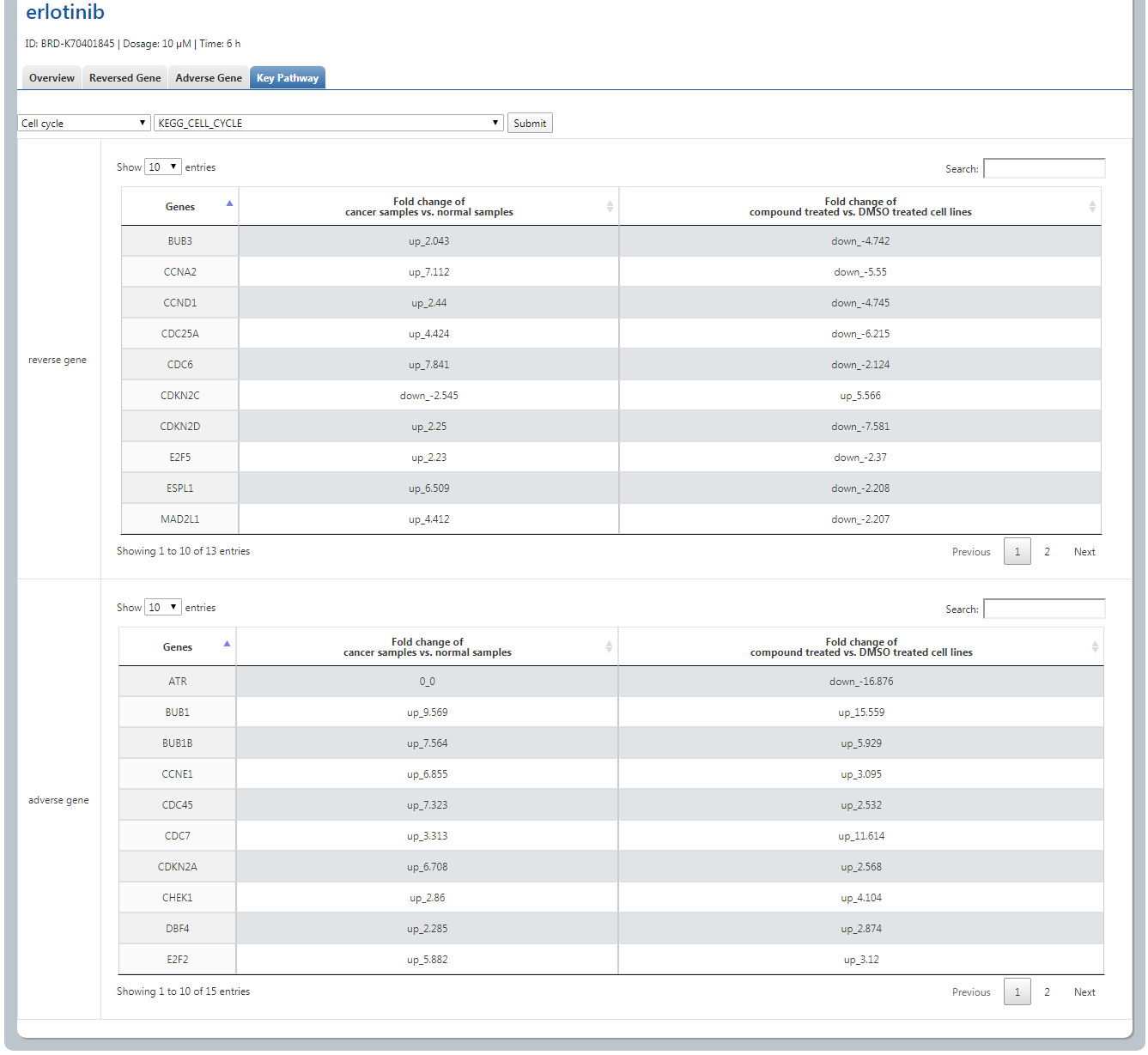
2. Personalized Medicine
Three options are provided.
(1) Enter Patient Matched Cell Line
When user enters cell line name, it will match the relative cancer samples of the same cancer type according to mutation similarity score by choosing “subTCGA vs. CMap (specific mutaion)” or “subTCGA vs. CMap (all mutation)”. We also provide an option of “TCGA vs. CMap” in case of user needs.
By clicking Detail for detail analyses information.
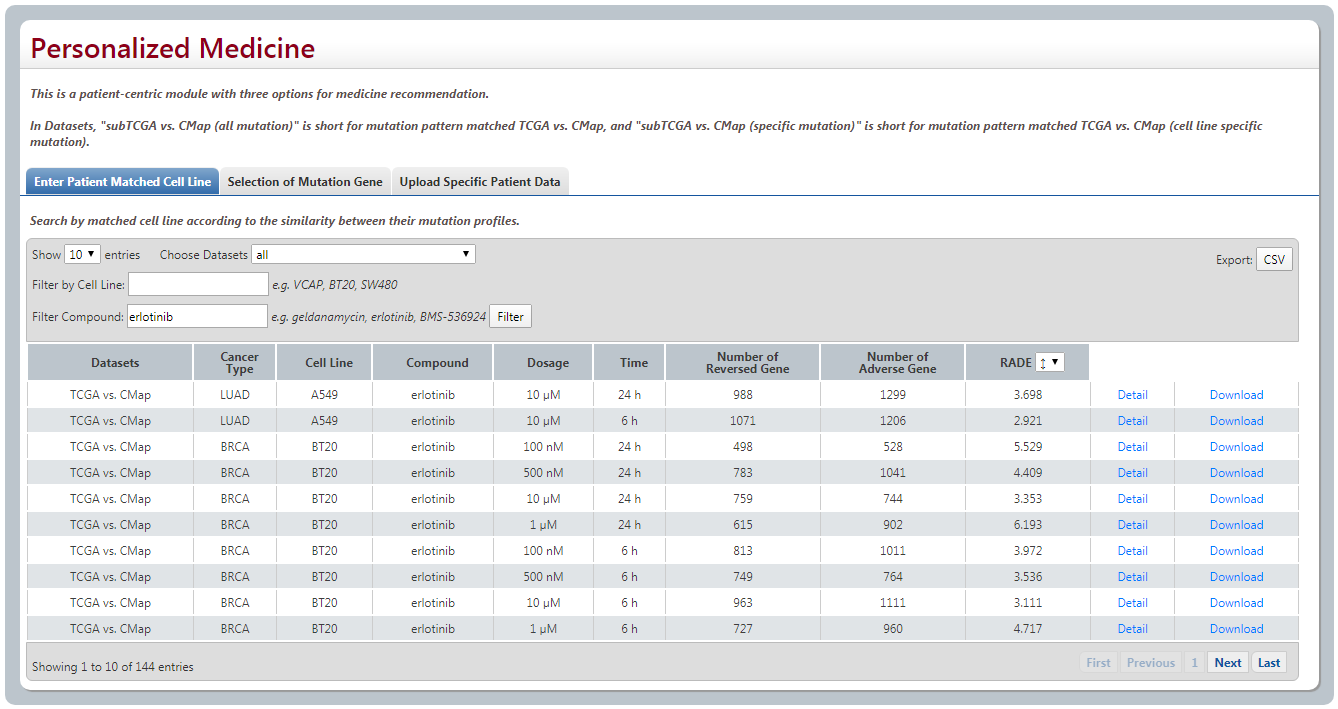
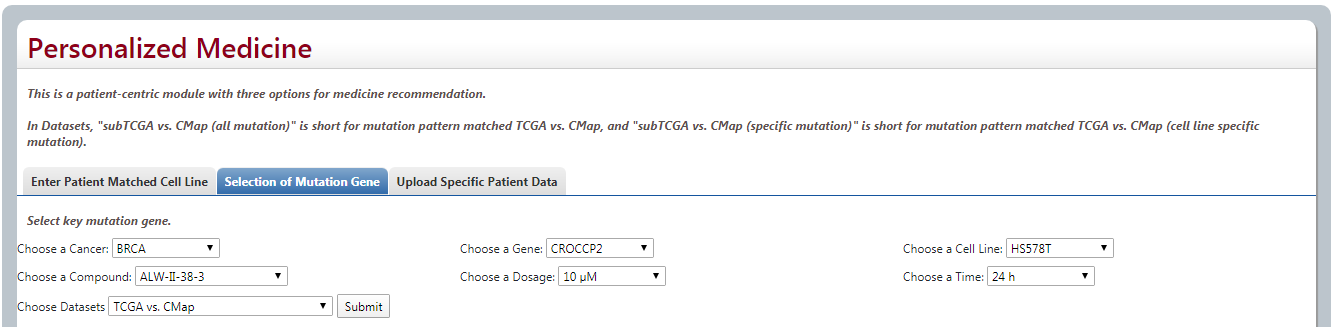
(2) Selection of Mutation Gene
For each cancer type, we provide a gene list in which each gene has at least 5% mutation rate in cancer samples of TCGA. Users could know the differences of drug response by patients with vs. without a specific mutation. After selecting cancer type, specific mutation gene, cell line, compound, dosage, time and datasets data orderly, it will list both the reversed effect and adverse effect of this drug on cancer patients with vs. without the mutation. Also User could download through 3 ways.
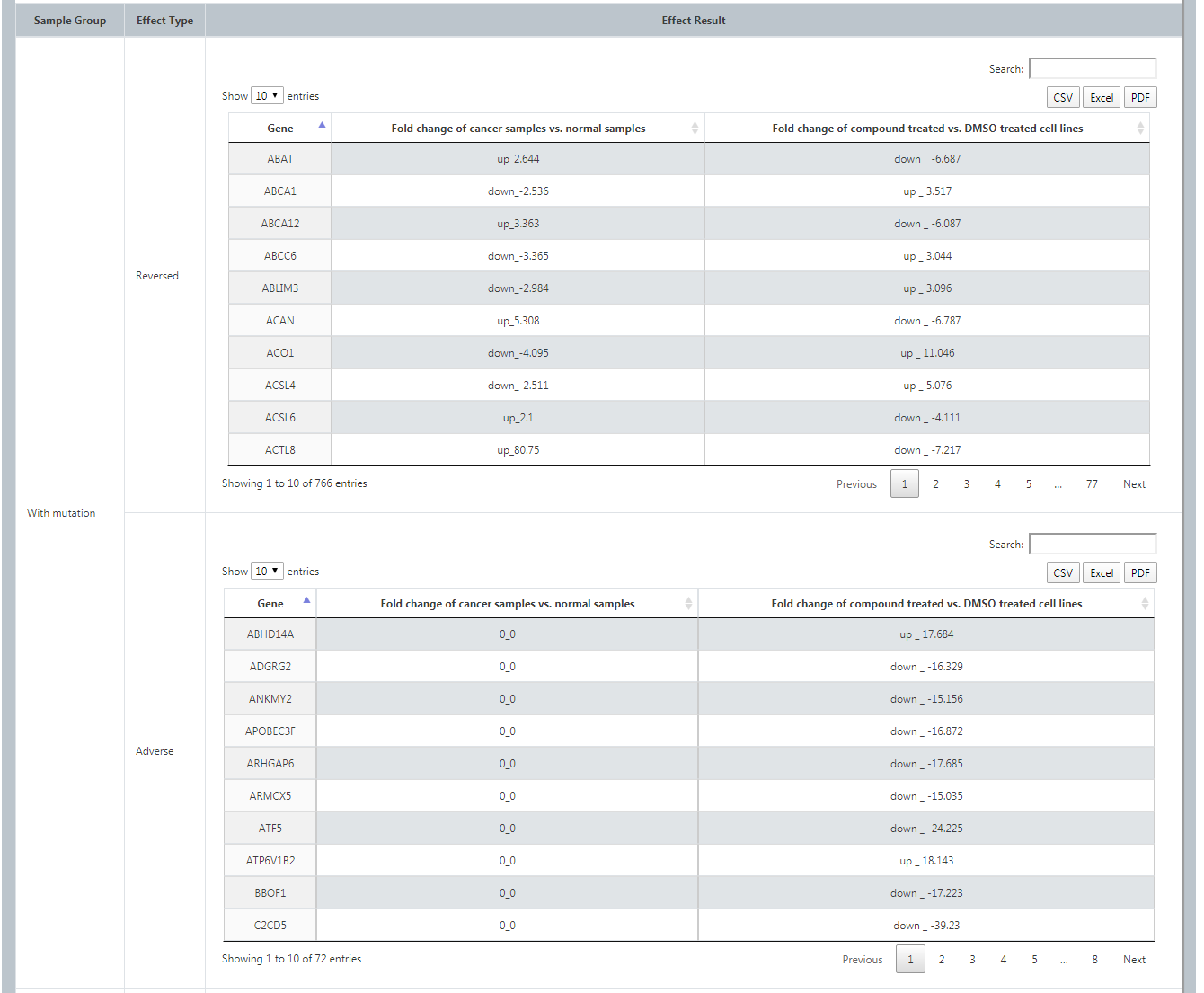
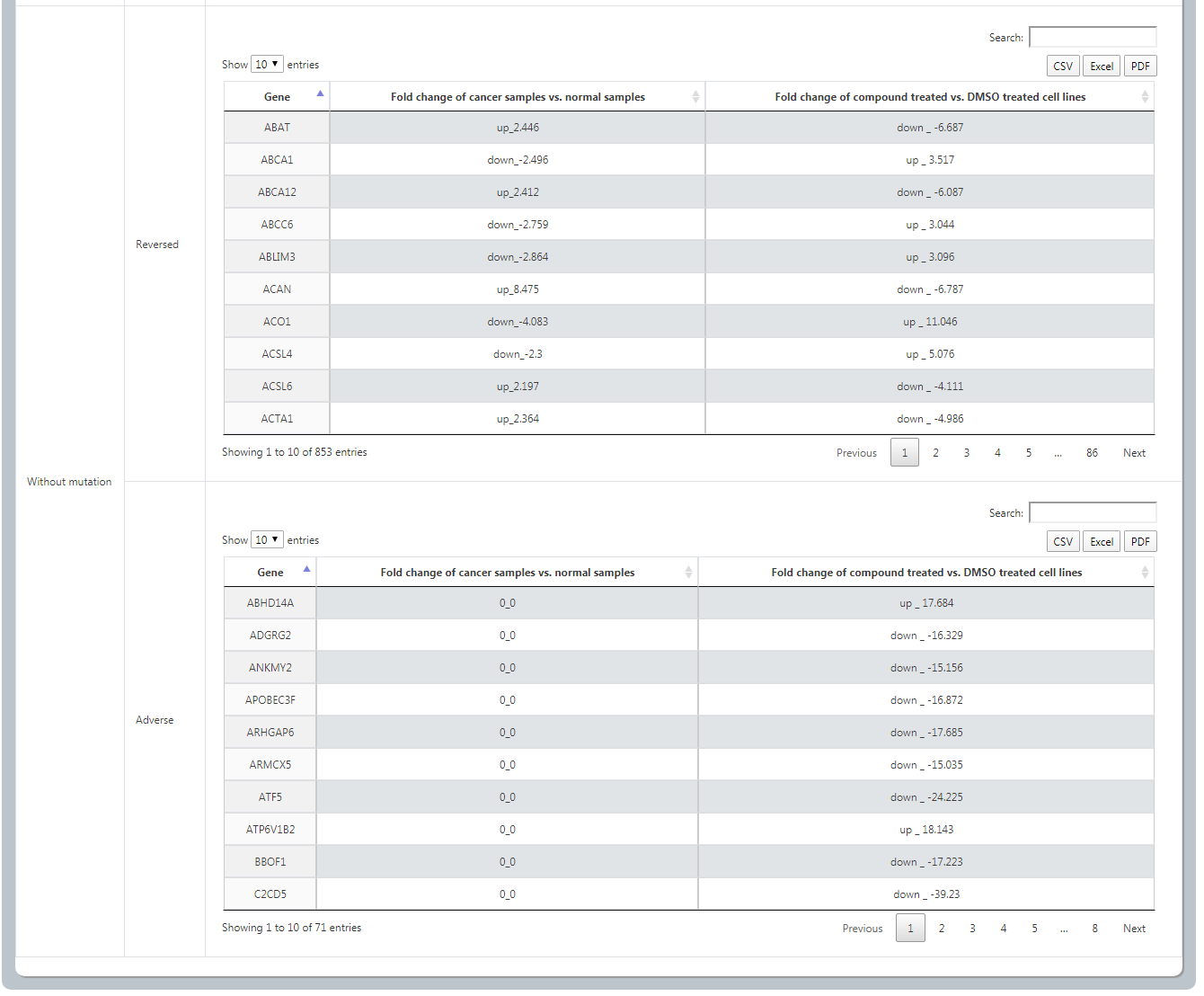
(3) Upload Specific Patient Data
Furthermore, user could also upload their own files about gene mutation profile of cancer sample for analysis. We provide an example file. After selecting this file, choosing the cancer type and then clicking uploading, the best matched cell line will be shown, following with the response that all the compounds on this cell line.
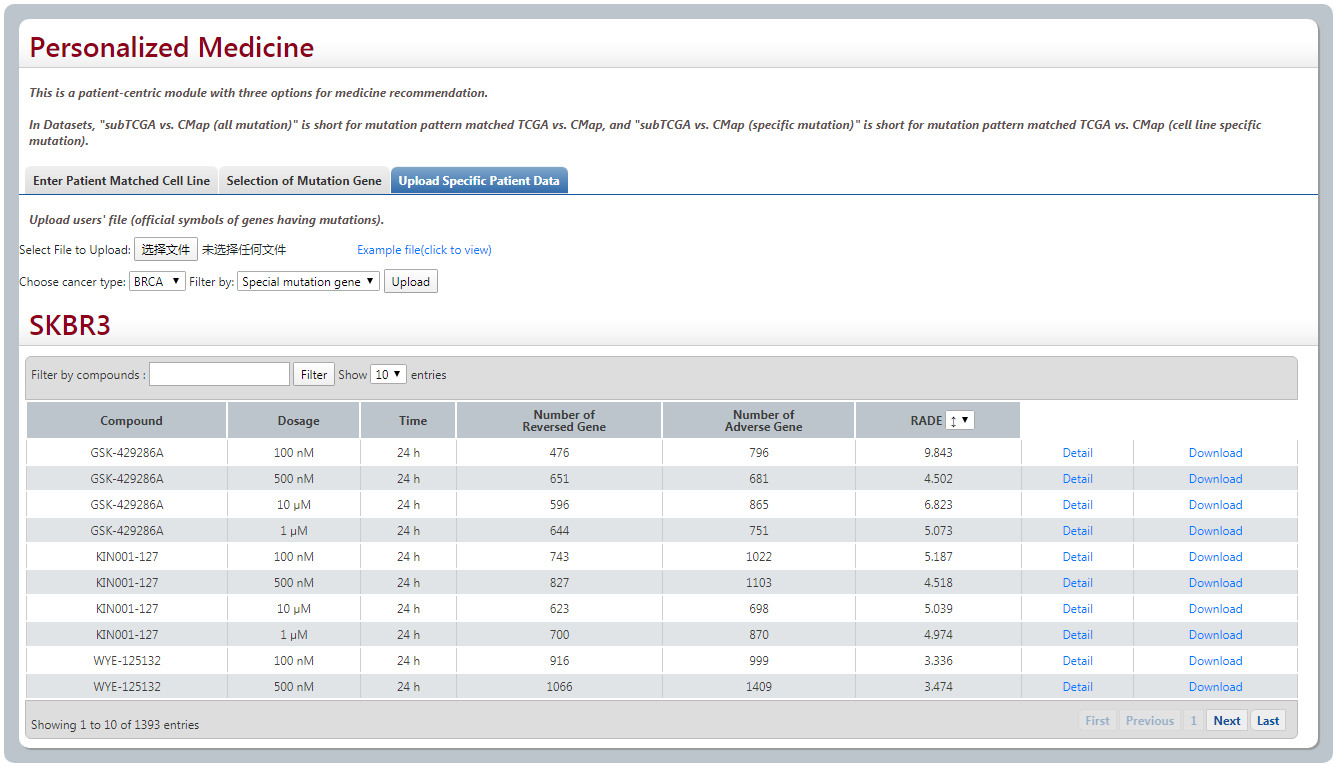
3. Drug combination
In this module, three options are available.
(1) 397 FDA approved drug pairs
When you select Drug A, Drug B and cell line, Two Venn Plots are provided to give the information about the intersection and different set of reserved genes and adverse genes that these two drugs affect, respectively. Besides, the detailed information are also shown as following.
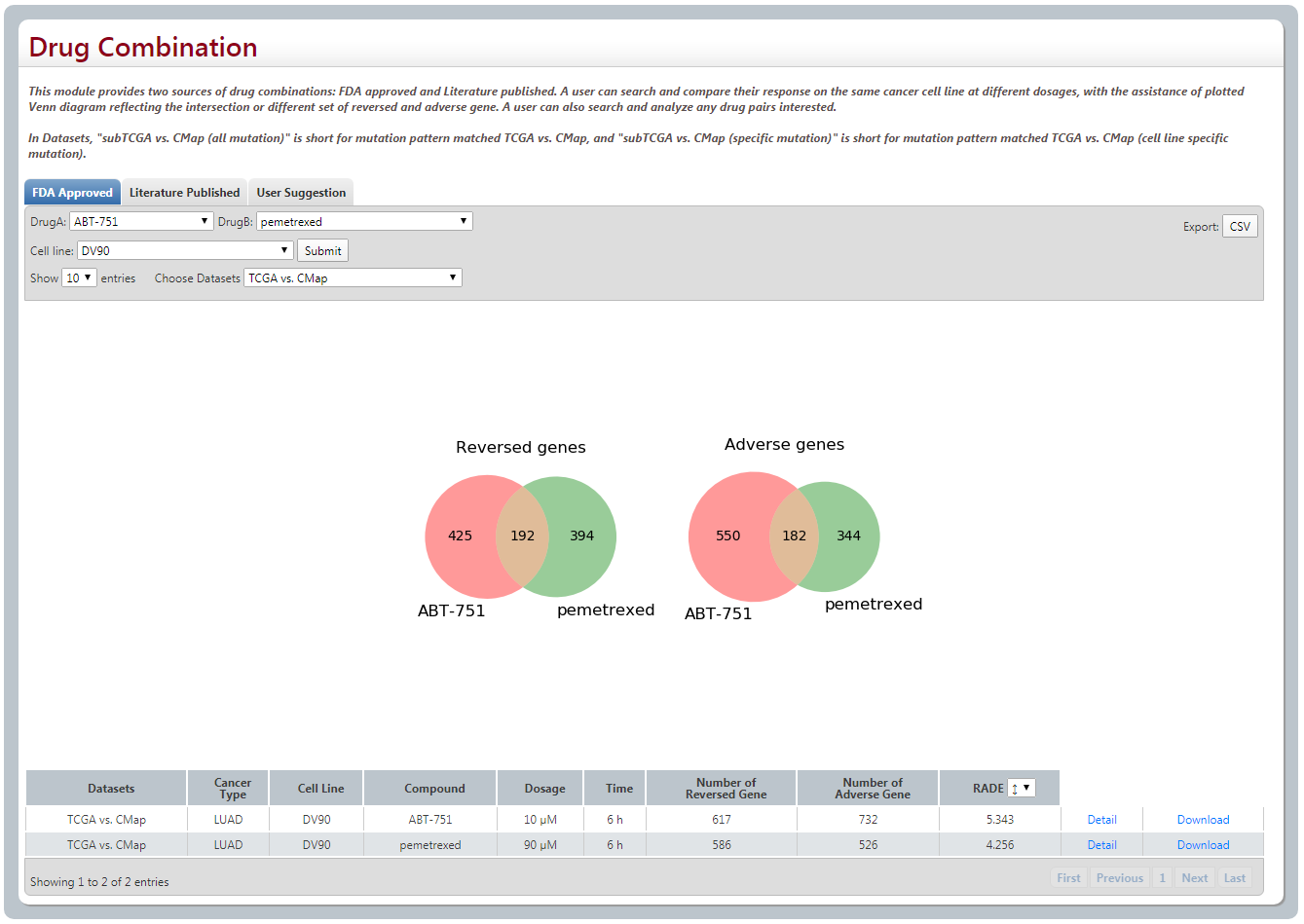
(2) 112 Literature published drug pairs. Similar with (1).
(3) User suggestion. User could search their interested drug pairs (in the download page, user could download all the compound we used in the database) for analysis.
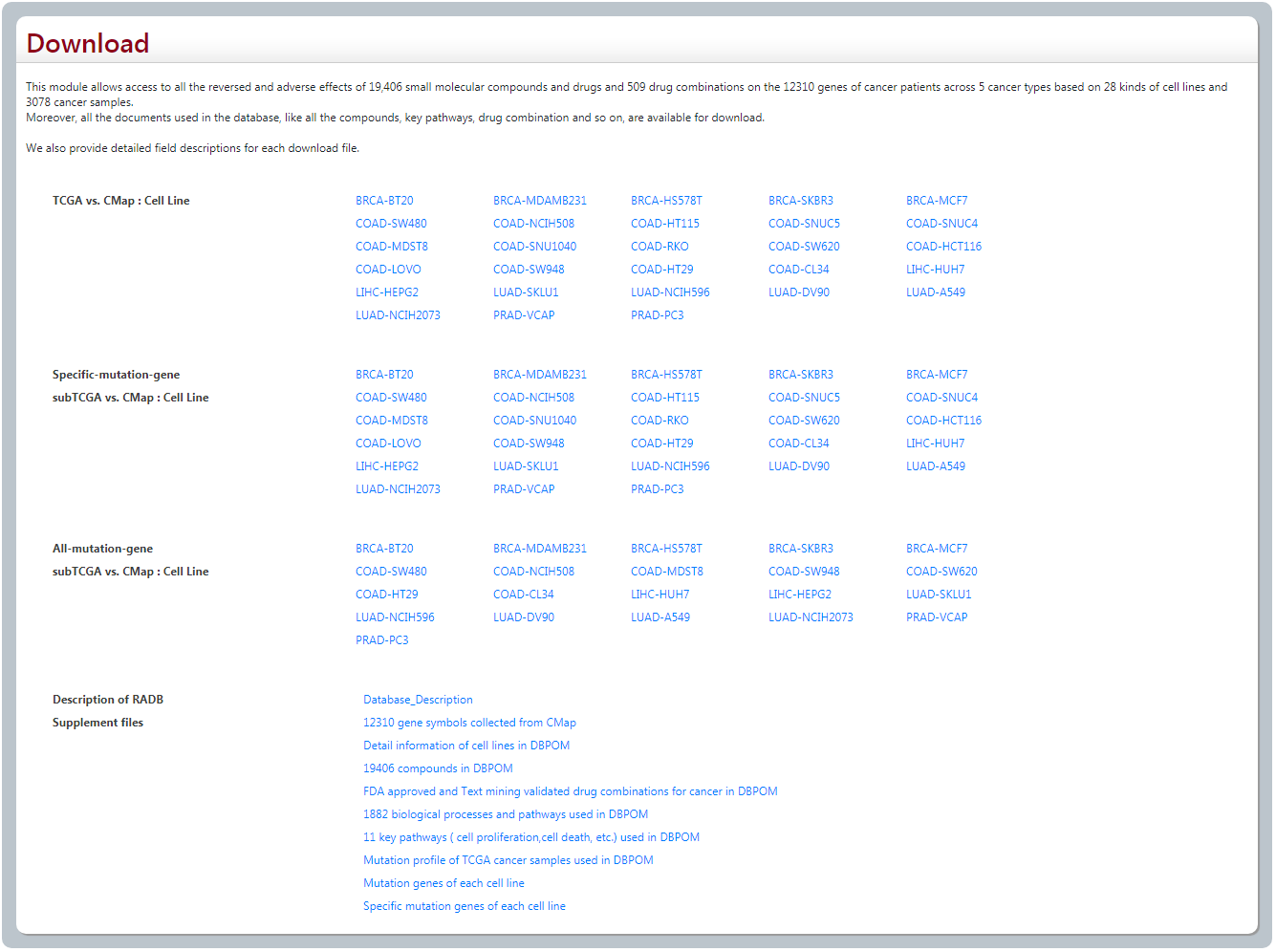
4. Download
User could download all the files and documents we used in DBPOM.